Visualizing MNIST Training Dynamics with Nomic Atlas¶
Nomic Atlas is a platform for interactively visualizing and exploring massive datasets. It automates the creation of embeddings and 2D coordinate projections using UMAP.
Nomic Atlas automatically generates embeddings for your data and allows you to explore large datasets in a web browser. Atlas provides:
- In-browser analysis of your UMAP data with the Atlas Analyst
- Vector search over your UMAP data using the Nomic API
- Interactive features like zooming, recoloring, searching, and filtering in the Nomic Atlas data map
- Scalability for millions of data points
- Rich information display on hover
- Shareable UMAPs via URL links to your embeddings and data maps in Atlas
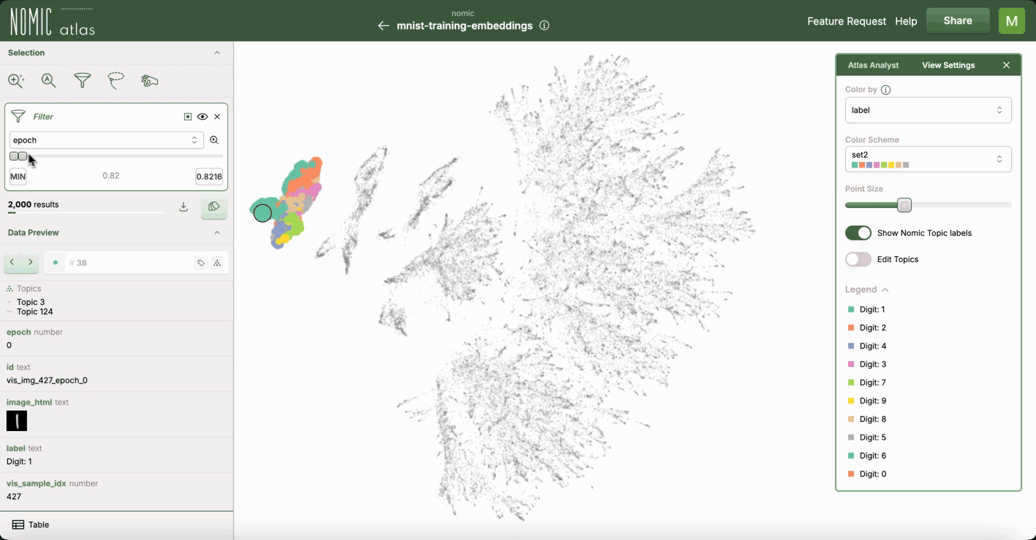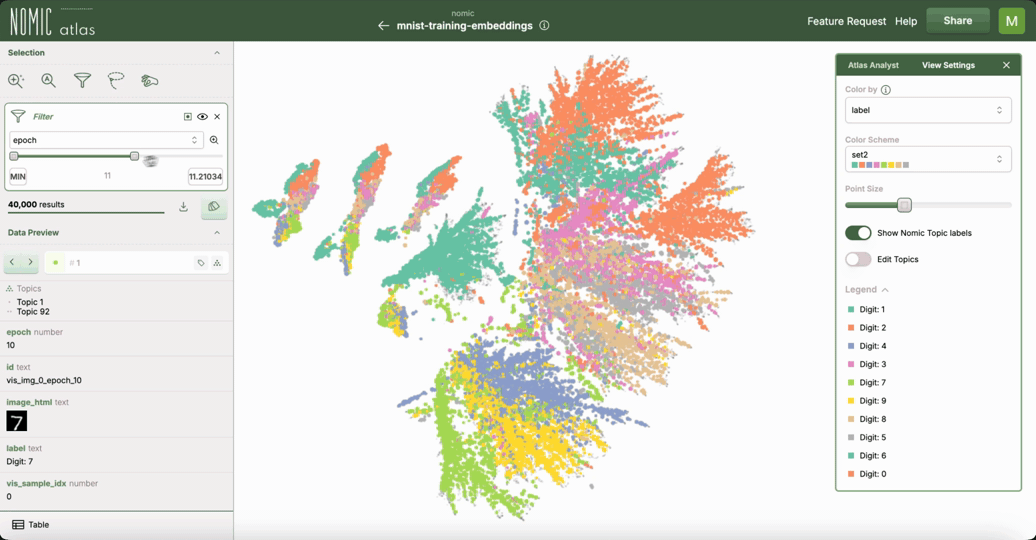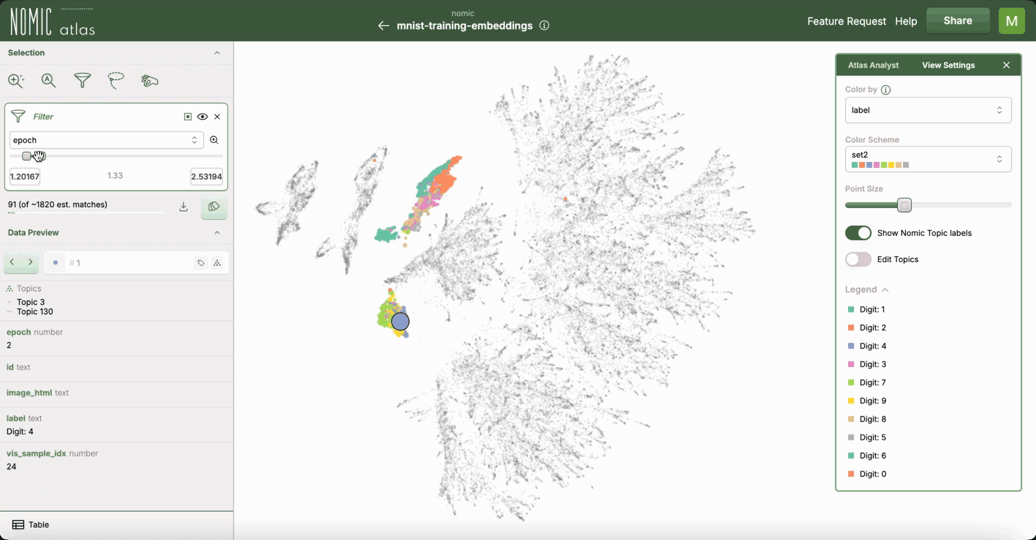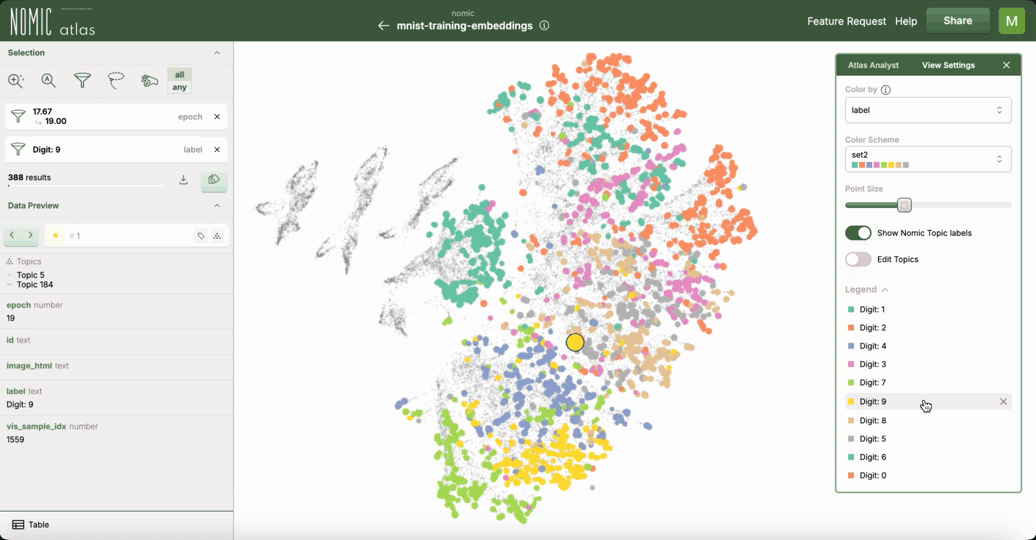
This example demonstrates how to use Nomic Atlas to visualize the training dynamics of your neural network using embeddings and UMAP.
Setup¶
- Get the python package with
pip install nomic - Get a Nomic API key here
- Run
nomic login nk-...in a terminal window or use the following code:
import nomic nomic.login('nk-...')
Download Example Data¶
import torch import torch.nn as nn import torch.optim as optim import torchvision import torchvision.transforms as transforms from torch.utils.data import DataLoader, Subset import numpy as np import time from PIL import Image import base64 import io
NUM_EPOCHS = 20 LEARNING_RATE = 3e-6 BATCH_SIZE = 128 NUM_VIS_SAMPLES = 2000 EMBEDDING_DIM = 128 ATLAS_DATASET_NAME = "mnist_training_embeddings" device = torch.device("cuda" if torch.cuda.is_available() else "cpu") print(f"Using device: {device}\n")
def tensor_to_html(tensor):
"""Helper function to convert image tensors to HTML for rendering in Nomic Atlas"""
# Denormalize the image
img = torch.clamp(tensor.clone().detach().cpu().squeeze(0) * 0.3081 + 0.1307, 0, 1)
img_pil = Image.fromarray((img.numpy() * 255).astype('uint8'), mode='L')
buffered = io.BytesIO()
img_pil.save(buffered, format="PNG")
img_str = base64.b64encode(buffered.getvalue()).decode()
return f''
class MNIST_CNN(nn.Module): def init(self, embedding_dim=128): super(MNIST_CNN, self).init() self.conv1 = nn.Conv2d(1, 32, kernel_size=3, padding=1) self.relu1 = nn.ReLU() self.pool1 = nn.MaxPool2d(kernel_size=2, stride=2) self.conv2 = nn.Conv2d(32, 64, kernel_size=3, padding=1) self.relu2 = nn.ReLU() self.pool2 = nn.MaxPool2d(kernel_size=2, stride=2) self.flatten = nn.Flatten() self.fc1 = nn.Linear(64 * 7 * 7, embedding_dim) self.relu3 = nn.ReLU() self.fc2 = nn.Linear(embedding_dim, 10)
def forward(self, x):
x = self.pool1(self.relu1(self.conv1(x)))
x = self.pool2(self.relu2(self.conv2(x)))
x = self.flatten(x)
embeddings = self.relu3(self.fc1(x))
output = self.fc2(embeddings)
return output, embeddings
transform = transforms.Compose([ transforms.ToTensor(), transforms.Normalize((0.1307,), (0.3081,)) ])
train_dataset = torchvision.datasets.MNIST(root='./data', train=True, download=True, transform=transform) test_dataset = torchvision.datasets.MNIST(root='./data', train=False, download=True, transform=transform)
persistent_workers_flag = True if device.type not in ['mps', 'cpu'] else False num_workers_val = 2 if persistent_workers_flag else 0 train_loader = DataLoader(train_dataset, batch_size=BATCH_SIZE, shuffle=True, num_workers=num_workers_val, persistent_workers=persistent_workers_flag if num_workers_val > 0 else False) vis_indices = list(range(NUM_VIS_SAMPLES)) vis_subset = Subset(test_dataset, vis_indices) test_loader_for_vis = DataLoader(vis_subset, batch_size=BATCH_SIZE, shuffle=False, num_workers=num_workers_val, persistent_workers=persistent_workers_flag if num_workers_val > 0 else False) print(f"Training on {len(train_dataset)} samples, visualizing {NUM_VIS_SAMPLES} test samples per epoch.\n")
Collect Embeddings During Training¶
model = MNIST_CNN(embedding_dim=EMBEDDING_DIM).to(device) criterion = nn.CrossEntropyLoss() optimizer = optim.Adam(model.parameters(), lr=LEARNING_RATE) all_embeddings_list = [] all_metadata_list = [] all_images_html = [] overall_start_time = time.time() for epoch in range(NUM_EPOCHS): epoch_start_time = time.time() model.train() running_loss = 0.0 for batch_idx, (data, target) in enumerate(train_loader): data, target = data.to(device), target.to(device) optimizer.zero_grad() outputs, _ = model(data) loss = criterion(outputs, target) loss.backward() optimizer.step() running_loss += loss.item() if (batch_idx + 1) % 200 == 0: print(f'Epoch [{epoch+1}/{NUM_EPOCHS}], Batch [{batch_idx+1}/{len(train_loader)}], Avg Loss: {running_loss / 200:.4f}') running_loss = 0.0 print(f"Epoch {epoch+1}/{NUM_EPOCHS} training finished in {time.time() - epoch_start_time:.2f}s.\n") model.eval() vis_samples_collected_this_epoch = 0 image_offset_in_vis_subset = 0 with torch.no_grad(): for data, target in test_loader_for_vis: data, target = data.to(device), target.to(device) , embeddings_batch = model(data) for i in range(embeddings_batch.size(0)): original_idx_in_subset = image_offset_in_vis_subset + i if original_idx_in_subset >= NUM_VIS_SAMPLES: continue all_embeddings_list.append(embeddings_batch[i].cpu().numpy()) img_html = tensor_to_html(data[i]) all_images_html.append(img_html) all_metadata_list.append({ 'id': f'vis_img{original_idx_in_subset}epoch{epoch}', 'epoch': epoch, 'label': f'Digit: {target[i].item()}', 'vis_sample_idx': original_idx_in_subset, 'image_html': img_html }) vis_samples_collected_this_epoch += 1 image_offset_in_vis_subset += embeddings_batch.size(0) if vis_samples_collected_this_epoch >= NUM_VIS_SAMPLES: break print(f"Collected {vis_samples_collected_this_epoch} embeddings for visualization in epoch {epoch+1}.\n") total_script_time = time.time() - overall_start_time print(f"Total training and embedding extraction time: {total_script_time:.2f}s\n")
Create Atlas Dataset¶
from nomic import AtlasDataset dataset = AtlasDataset("mnist-training-embeddings")
Upload to Atlas¶
dataset.add_data(data=all_metadata_list, embeddings=np.array(all_embeddings_list))
Create Data Map¶
We specify the text field from df as the field to create embeddings from. We choose some standard UMAP parameters as well.
dataset.create_index(projection='umap', topic_model=False)
Your map will be available in your Atlas Dashboard.